Why this unit
This unit links connectome graph analysis to AI-relevant hypothesis generation.
Learning goals
- Build motif-search workflows from hypothesis to interpretation.
- Evaluate statistical and computational limits of graph-based inference.
Core technical anchors
- Null-model comparisons and multiple-comparison controls.
- Subgraph isomorphism complexity and tooling tradeoffs.
- Cross-dataset comparability constraints.
Method deep dive: motif-analysis pipeline
- Hypothesis formalization:
Convert biological intuition into graph constraints and measurable outputs.
- Query implementation:
Encode motifs in a query language and validate on synthetic control graphs.
- Search execution:
Run distributed motif scans with resource/latency monitoring.
- Statistical testing:
Compare observed counts to null ensembles and apply multiplicity corrections.
- Biological interpretation:
Connect motif enrichments to plausible circuit mechanisms while stating uncertainty.
Model and inference considerations
- Representation choices:
Directed vs undirected edges, weighted vs binary synapses, multigraph encoding.
- Null-model families:
Degree-preserving rewires, spatially constrained rewires, cell-type-stratified controls.
- Scalability tradeoffs:
Exact subgraph isomorphism vs approximate search and candidate pruning.
- Reproducibility:
Pin dataset versions, query code revisions, and random seeds.
Quantitative QA checkpoints
- Query correctness on toy graphs with known motif counts.
- Runtime and memory benchmarks by motif size/complexity.
- Sensitivity analysis across alternate null-model definitions.
- Stability checks across reconstruction versions and proofreading updates.
Frequent failure modes
- Post-hoc hypothesis selection:
Separate exploratory and confirmatory analyses.
- Null-model mismatch:
Ensure nulls preserve the structural constraints relevant to the claim.
- Cross-dataset overgeneralization:
Treat species/region-specific findings as context-bound unless replicated.
- Toolchain opacity:
Require auditable query scripts and logged execution parameters.
Practical workflow
- Define biologically grounded motif hypotheses.
- Translate hypotheses into executable graph queries.
- Run searches across selected connectome datasets.
- Compare motif prevalence with null models and controls.
- Interpret results with explicit statistical and dataset assumptions.
Visual training set (draft)
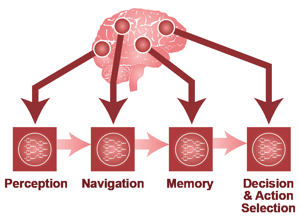
Techtalk S10: motivation question linking natural and artificial intelligence.

Techtalk S11: brain-data framing for analysis context.

Techtalk S12: reverse-engineering analogy for computational decomposition.

Techtalk S13: NeuroAI pipeline framing.
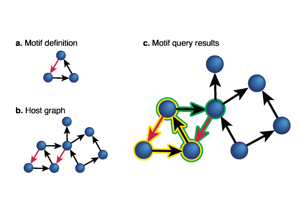
Techtalk S24: subgraph motif-search concept.

Techtalk S26: query-language/tooling transition.
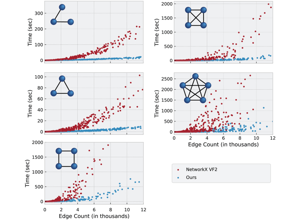
Techtalk S31: subgraph-isomorphism algorithm context.
Techtalk S32: performance-benchmark context.

Techtalk S33: throughput/scale context (fallback for missing S34 extraction).

Techtalk S39: atlas-scan hypothesis example.
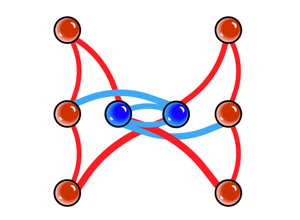
Techtalk S42: DotMotif syntax and query expression example.
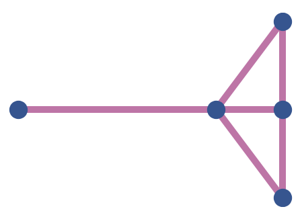
Techtalk S44: developmental motif-comparison context.

Module13 L3 S03: project-overview context.

Module13 L3 S11: data-growth and scale context.

Module13 L3 S14: processing-comparison context.

Module13 L3 S20: connectivity-estimation context.

Module13 L3 S24: classification/model context.

Module13 L3 S29: late-stage synthesis context.

Module13 L3 S37: application-stage context.
Attribution: NeuroAI and outreach source decks from the extraction package. Historical figures (including 2021 techtalk materials) are used for technical context; interpret benchmark claims as historical unless independently revalidated.
Discussion prompts
- What evidence is needed before treating a motif finding as functionally meaningful?
- Which null models are most defensible for a given connectome dataset?
- Where do computational constraints shape scientific conclusions in this workflow?
Course links
Quick activity
Define one motif hypothesis, one null model, and one success criterion you would use before interpreting results.
Draft lecture deck



















