Why this unit
Glia are central to reliable annotation and interpretation, not background objects.
Learning goals
- Recognize astrocytes, microglia, and oligodendrocyte cues.
- Reduce glia-neuron boundary errors during proofreading.
Core technical anchors
- Glia-specific ambiguity patterns in dense neuropil.
- Myelin context cues for oligodendrocyte interpretation.
- Reusable glia recognition checklist.
Method deep dive: glia identification workflow
- Candidate detection:
Mark processes with non-neuronal morphology and atypical organelle distribution.
- Context enrichment:
Inspect vascular adjacency, myelin relationships, and neighboring synaptic density.
- Cell-class discrimination:
Compare astrocyte-like branching, microglial surveillance morphology, and oligodendrocyte/myelin patterns.
- Temporal/section continuity:
Follow process across slices to avoid single-plane misclassification.
- Adjudication:
Escalate low-confidence cases into a shared glia review queue.
Quantitative QA checkpoints
- Glia-vs-neuron boundary error rate on validation subset.
- Class-specific agreement (astrocyte, microglia, oligodendrocyte).
- Rate of unresolved glia labels after second-pass review.
- Impact of corrected glia labels on neuronal graph statistics.
Frequent failure modes
- Astrocyte-neurite confusion in dense neuropil:
Require neighborhood and boundary-context confirmation.
- Over-calling microglia based on fragmentary appearance:
Use multi-slice morphology before class assignment.
- Oligodendrocyte/myelin ambiguity:
Trace sheath context and nearby axonal relationships.
- Ignoring glia in proofreading priorities:
Include glia corrections in high-impact QC queues.
Visual training set

RIV-GLIA S01: opening context visual for glia-focused proofreading.

RIV-GLIA S03: astrocyte-related morphology and synaptic neighborhood context.
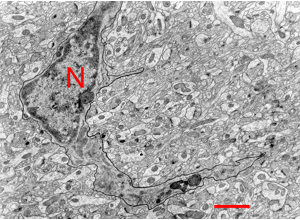
RIV-GLIA S09: microglia recognition cues in local structural context.

RIV-GLIA S15: oligodendrocyte-focused morphology/reconstruction cue.

RIV-GLIA S16: myelin-related context for glia interpretation.
Attribution: Pat Rivlin training materials (MICrONS proofreading deck). Two manifest-listed IDs (`S02`, `S07`) were not present in extracted thumbnails and are pending recovery.
Course links
Practical workflow
- Identify candidate glial morphology in local context.
- Compare against neuronal look-alikes in neighboring slices.
- Confirm with vascular/myelin/synaptic adjacency cues.
- Record class decision and uncertainty for review.
Discussion prompts
- Which glia-neuron ambiguities are most error-prone in your data?
- What minimum evidence should be required before final glia labels?
Quick activity
Review one glia image and list two features that distinguish it from a neuronal process in the same neighborhood.
Content library references
Teaching slide deck
Evidence pack: papers and datasets
This unit is anchored to canonical papers and datasets used in connectomics practice. Use these as required preparation before activities.
Key papers
Key datasets
Competency checks
- Separate neuron and glia boundaries in mixed patches with audit notes.
- Identify high-risk glia-neuron confusion zones for targeted QC.
Capability development brief
Capability target: Identify major glial classes and prevent glia-neuron boundary errors in reconstruction workflows.
Required expertise
- Glia biologist (cell-class and functional context)
- EM proofreader (boundary and myelin interpretation)
- QC lead (error auditing and process controls)
Core concepts to teach
- Glia class cues: Distinctive ultrastructural signatures for astrocytic, oligodendroglial, and microglial profiles.
- Myelin context: Interpreting sheaths and associated processes to avoid identity leakage.
- Boundary integrity: Maintaining consistent neuron-glia separation across long trajectories.
Studio activity
Glia Boundary Audit - Detect and correct glia-neuron confusions in realistic proofreading samples.
Audit mixed patches and classify all uncertain boundaries by error type.
- Identify glial signatures and map candidate boundaries.
- Tag high-risk boundary zones for targeted review.
- Propose correction order based on downstream impact.
Expected outputs:
- Boundary audit table
- Prioritized correction queue
Assessment artifacts
- Glia identification quick-reference guide.
- Boundary error audit with prioritized correction rules.





